Publication
909
ACS Catal., 10, 7843-7856, 2020
DOI:10.1021/acscatal.0c01920
|
|
|
|
|
|
|
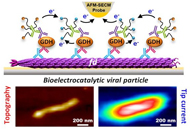
Probing the Enzymatic Activity of Individual Biocatalytic fd-Viral Particles by Electrochemical-Atomic Force Microscopy |
|
|
|
Telmo Manuel Oliveira Paiva, Kristian Torbensen, Anisha Patel, Agnès Anne, Arnaud Chovin, Christophe Demaille, Laure Bataille, and Thierry Michon
Université de Paris, Laboratoire d’Electrochimie Moléculaire LEM, CNRS, F-75013 Paris, France
Université de Bordeaux, Biologie du Fruit et Pathologie, INRA, F-33140 Villenave d’Ornon, France
Surface-immobilized fd bacteriophage particles are used as scaffolds to coassemble the redox enzyme quinoprotein glucose dehydrogenase, PQQ-GDH, and its cosubstrate, PEG-tethered ferrocene. Individual decorated fd phages are visualized and simultaneously functionally interrogated by Mt/AFM-SECM microscopy, an in-situ local probe correlative imaging technique, combining atomic force (AFM) and electrochemical (SECM) microscopy in a mediator tethered (Mt) configuration. The statistical distribution of catalytic activity across the fd population is resolved, and the correlation between the functional properties of the phages and their actual dimensions assessed. Moreover, achievement of sub-particle resolution allows the enzymatic activity of individual viruses to be spatially mapped, revealing a highly active region located in the middle of the filamentous fd-scaffold. Quantitative modeling shows that this “catalytic hot-spot” arises from the interplay between charge transport by electron hopping between ferrocene moieties along the viral particles, and enzymatic catalysis. The developed model also enables complete analysis of GDH kinetics at the single bioscaffold scale, revealing differences in the functional behavior of the biocatalytic viral particles when addressed at the ensemble or at the single particle scale. |

